My name is Katie Jo Gelasco and I am a junior pursuing my dual degree in music and biochemistry. While I have loved playing the cello since the young age of six, my other passion lies in biochemistry. After graduation, I hope to get my Phd in biochemistry or cancer biology and develop a career in medical research. Currently, I am very involved in the College of Music, where I take weekly cello lessons and am a member of the University Symphony Orchestra. We have recently returned from our first international tour to the Dominican Republic. I am on the executive board of the Gamma Beta chapter of the national chemistry fraternity, Alpha Chi Sigma. As the Alumni Secretary, I write a quarterly newsletter and organize events that will increase the professional development of our members. I also run and organize our chapter’s website.

Currently, I am an undergraduate member of the Li lab in the Kasha Institute of Molecular Biophysics, where we focus on exploring the structure and function of CRISPR-Cas9 proteins for use in broader application. Discovered by Jennifer Doudna and Emmanuelle Charpentier in 2011, these versatile proteins have been the subject of thousands of publications and are an exciting, and even controversial approach to gene editing. This revolutionary technology could lead to a future of modified plants, personalized gene therapies, and even treatments for diseases such as HIV and cancer. While most labs focus on application of CRISPR-Cas9 proteins, we are one of the few that engineers these proteins specifically to study the function and biochemical mechanism during cleaving of DNA.
CRISPR is a heritable, immune response found in bacteria that defends against bacteriophages. When a phage infects a cell, the cell can capture small pieces of the virus’ DNA and insert it into its CRISPR regions. After these sequences are transcribed into CRISPR RNA, they are combined with an endoribonuclease Cas (CRISPR-associated) protein, and work to find the matching DNA pieces in the virus. The guide RNA recognizes the target DNA via its PAM (protospacer adjacent motif) and guides the Cas protein to cleave the DNA, inactivating the virus. Researchers can use Cas proteins from this immune response to edit nucleic acids, such as DNA, by designing a single-guide (sgRNA) that binds to Cas9 proteins to target and destroy virtually any gene they design it for. These proteins have high specificity in detecting a target DNA sequence, thanks to PAM recognition. Researchers can also replace the cut DNA with template DNA using directed homology repair to edit DNA sequences, making it a much easier and more efficient way to genetically edit in organisms.
My project will focus on engineering the phosphate lock residues of the Cas9 protein in Streptococcus pyogenes. A previous publication from our lab explored the effects of manipulating the phosphate lock loop residues (amino acid sequence) on the catalytic efficiency of Cas9 in the novelty thermophilic bacteria, Acidothermus cellulolyticus. The phosphate lock loop on the Cas9 protein interacts with the phosphate group of the DNA target strand near the PAM sequence. AceCas9’s PAM sequence is 5’-NNNCC-3’, and this PAM sequence is unique in every species of Cas9 protein. It was supported by previous studies that mutational manipulation of the phosphate lock loop can ease the PAM requirement of AceCas9.
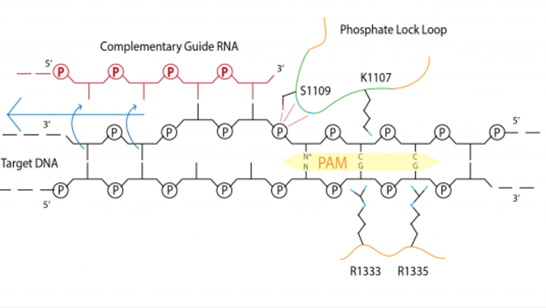
My project will focus on engineering the SpyCas9 found in Streptococcus pyogenes, which is much more of interest to the biosciences community, since SpyCas9 activity has been shown to successfully edit human genomes in human cells. This connection between PAM specificity and the phosphate lock loop has yet to have been explored in SpyCas9. CRISPR-Cas9 gene editing requires the use of a PAM sequence to orient the Cas9 protein for it to cleave the correct target. This PAM sequence is unique to every bacteria with SpyCas9’s being 3’NGG, which means it must find and recognize this nucleotide base sequence before cleaving the DNA. This signifies that there is a finite amount of targetable DNA sequences. This is restrictive towards broader application, and researchers are searching for either a PAM-free endonuclease or mechanisms to relax the PAM requirement. So while a guide RNA to target a certain gene for editing can be engineered by researchers, it will need to recognize the specific PAM sequence before ever binding to the target strand. My project will feature a directed protein evolution of SpyCas9 phosphate lock loop in order to ease the PAM requirement. I will also determine if this altered Cas9 protein has increased the catalytic efficiency of the enzyme.
This grant will support my work in the lab this summer, including funding a SpyCas9 phosphate lock loop library. This will provide variants of the protein at the phosphate lock loop position that allow for rescue of enzyme (cleavage of lethal gene) activity in unfavorable conditions. Then, we will select for the variants that allow for survival and evolve the protein to eventually relax the requirement for its PAM sequence.
I am excited for the opportunities this grant has provided me and I am thrilled to begin work on my project this summer.