My name is Zehta Glover and I am a senior in the Department of Biological Science where I am majoring in Cell and Molecular Neuroscience. During my sophomore year, I began working in Dr. Hank W Bass’ research lab studying maize genetics. Over the last two years, I have grown intrigued by research and have since grown independence in the lab. This has encouraged me to begin an Honors in the Major Thesis (HITM), which will be funded by the IDEA grant. My time performing research has encouraged me to look into careers in a lab setting. While I’m not exactly sure what that entails at this point of my academic career, I am considering pursuing a PhD in neuroscience or entering the work force as a forensic or medical lab technician.

Lisa Scott Endowed Undergraduate Research Award winner
Now, less about me and a little bit more about my research. The research project for which I have been funded by the IDEA grant for is titled “Analysis of micrococcal nuclease properties relative to chromatin profiling assays in maize (Zea mays L.) and human.” In order to properly introduce what I will be doing, I think it is first important to explain why I will be performing this research.
Understanding gene regulation is a major goal in biology as it is vital for growth and
development of all organisms. While much research has been done to answer the unknowns of gene regulation, many questions remain unanswered. To fully understand gene regulation, we must understand chromatin structure, which includes transcription factor (TF) binding sites and their occupancy (reviewed by Tsompana and Buck, 2014). In Dr. Hank W Bass’ research lab, Dr. Bass and students have used an enzyme called micrococcal nuclease (MNase) to cut regions of maize DNA and align it to the maize genome to further analyze topics, including, gene regulation. However, the enzyme MNase has been critiqued for its possible bias to cut DNA at AT-rich regions (Allan et al., 2012).
My research project will analyze MNase’s sequence bias through control tests and a new methodology, in which I will be creating, called a gain-of-footprint assay. I will first be completing the control tests. For this, my first step would be to obtain high molecular weight naked DNA. In the lab, I have isolated maize DNA using the Qiagen Plant Maxi DNA kit. Using this kit, I was able to take maize earshoot tissue that was ground into a fine powder, and isolate the DNA through a series of disruptions and separations. Human DNA was generously donated by the J.H. Dennis Lab at FSU and by the Gillespie Lab at Univ. South Alabama. Below are images of the DNA in a 1% Agarose gel. Through these gels, I was able to see that I had high molecular weight naked DNA in at least most of my samples.
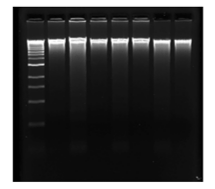
Now that I have collected total naked DNA for both maize and human, I will be working on the MNase digestion. This technique will be similar to that described as MOA-seq by Savadel et al., (2021). For this, I will prepare a series of MNase dilutions, allowing MNase to digest the DNA at different concentrations. Throughout this process I will observe the MNase digestions through Gel electrophoresis. Once I observe the desired Gel electrophoresis and reach the dilution that carries out the preferred light partial digest of DNA, I will make libraries of the DNA fragments and sequence and align them to the genome. For the gain-of-footprint assay, I will complete the same exact steps mentioned in the previous paragraph; however, before I begin the MNase digestion, I will insert a known DNA binding protein (CTCF) into the DNA. Then again, sequence and align the fragments from MNase digestions to the genome.

For my last step, I will characterize the MNase digestion patterns. Currently I am still working out the specific details of how I will analyze the results. However, I expect to see reads of small fragments centered around the binding sites of the known DNA binding protein. Then I will compare these footprints to the footprints I receive from the control tests. This comparison should allow me to conclude whether MNase has a bias to cut at AT-rich regions of DNA. Over the summer I hope to complete most, if not, all of the wet bench procedures and then begin working on the analysis. I’ve already made so much progress in the lab thus far, and am excited to spend my summer doing research!
Photo by National Cancer Institute on Unsplash
